Syndrome Surveillance Tools & SEAM
This project focuses on providing VA clinical research with methods and instruments for syndrome surveillance, on improving detection and mitigation of health problems in deployed veterans, and more specifically on the detection of medically unexplained syndromes (MUS) and emergent infectious diseases.
Developing tools to effectively use information extracted from VA clinical documents depends on expertise in information extraction, ontology development, symptom identification, predictive modeling, and interactive information visualization. Thus, execution of the work entailed by this project depends on the capacity to work across a range of clinical areas, and to engage a scientific team that brings together experts from a variety of disciplines including epidemiology, infectious disease, mental health, computer science, biomedical informatics, and statistics.
We were organized in four groups led by Dr. Stephane Meystre (information modeling and ontologies, and overall leadership), Dr. Scott Duvall (information extraction), Dr. Mollie Cummins (data mining), and Dr. Yarden Livnat (information visualization).
Semi-automated ontology development and management (SEAM) application: For the development of this application, we conducted an extensive review of existing methods and tools for terms, concepts, and relation discovery in unstructured text. We used the review findings to inform the development of the ontology management system. For the user interface, we based our developments on two ideas: card sorting and concept laddering. The former is a simple method to elicit categorization of concepts, giving the user functionalities to group ‘cards’ (automatically extracted terms and concepts in our case) into categories they can name. The latter is a method to organize concepts in hierarchies, giving users functionalities to organize concepts in taxonomies and create relations between them.
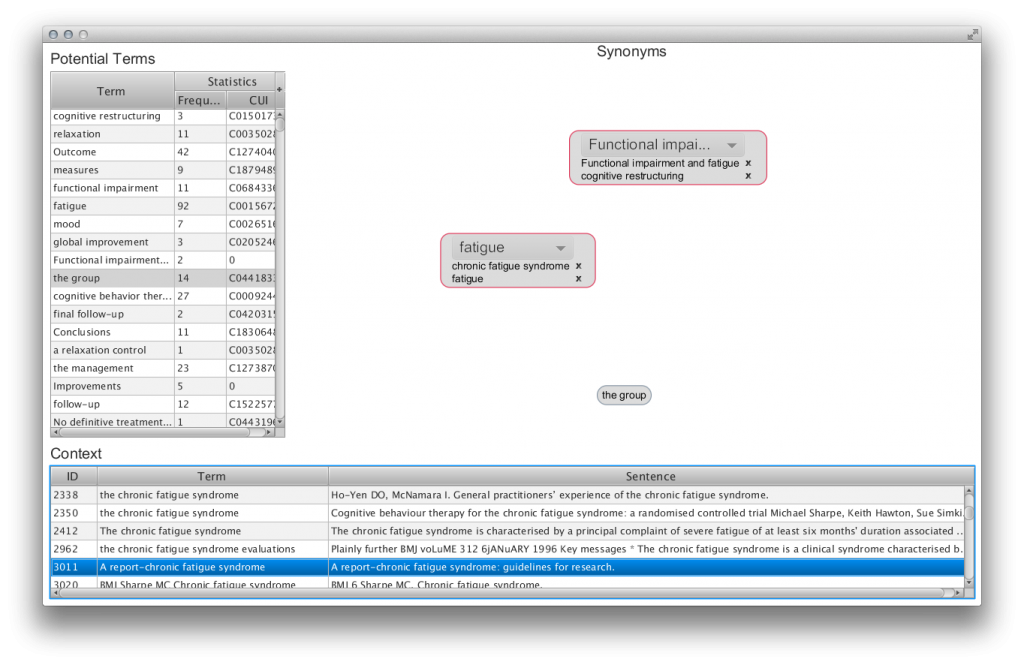
Publications:
- Doing-Harris, K., Livnat, Y., & Meystre, S. M. (2015). Automated concept and relationship extraction for the semi-automated ontology management (SEAM) system. Journal of Biomedical Semantics, 6(1), 1–15.
- Meystre, S. M., Doing-Harris, K., Boonsirisumpun, N., Potter, K., & Livnat, Y. (2013). Semi-Automated Ontology Development Systemfor Medically Unexplained Syndromes in the U.S. Veterans Population (pp. 1–1). AMIA Annu Symp Proc.
- Doing-Harris, K., Meystre, S. M., Samore, M., & Ceusters, W. (2013). Applying ontological realism to medically unexplained syndromes. Studies in Health Technology and Informatics, 192, 97–101.
